Diffusion-Based Counterfactual ECG Generation for Atrial Fibrillation Data Augmentation
Team
- E/20/069, Tharaka Dilshan, e20069@eng.pdn.ac.lk
- E/20/189, Nethmini Karunarathne, e20189@eng.pdn.ac.lk
Supervisors
- Dr. Vajira Thambawita, SimulaMet, Oslo, Norway
- Prof. Mary M. Maleckar, Tulane University / Simula Research Laboratory
- Dr. Roshan Ragel, University of Peradeniya
- Dr. Isuru Nawinne, University of Peradeniya
Collaborators
- Isuri Devindi, University of Maryland, College Park, USA
- Prof. Jørgen K. Kanters, University of Copenhagen, Denmark
Table of Contents
Introduction
Automated detection of atrial fibrillation (AFib) using deep learning requires large, balanced ECG datasets, yet clinical data remains scarce, imbalanced, and constrained by privacy regulations. We present a diffusion-based data augmentation pipeline that generates synthetic ECG segments by transforming existing recordings into counterfactual waveforms of the opposing class. The pipeline uses a partial-noise conditional denoising process with classifier-free guidance, operating on single-lead ECG signals. A content-style disentangled UNet architecture separates class-invariant morphology from class-discriminative rhythm features. A multi-stage plausibility post-validator enforces morphological and physiological constraints, retaining only waveforms that satisfy quality thresholds.
The augmented mixture achieves 95.05% accuracy and 98.60% AUROC, statistically equivalent to original-only training (95.63% accuracy, TOST p = 0.007, Δ = ±2%). Furthermore, none of the accepted counterfactuals are near-copies of training data (maximum correlation: 0.30), indicating the generated signals are novel and privacy-preserving.
This research is conducted as part of the EU-funded SEARCH Initiative through an international collaboration between the University of Peradeniya (Sri Lanka), SimulaMet (Norway), Tulane University (USA), and the University of Copenhagen (Denmark).
Related Works
Generative models for ECG synthesis have been explored using GANs, VAEs, and more recently, diffusion models.
| Method | Model | Counterfactual | Key Limitations |
|---|---|---|---|
| PGAN-ECG (Golany et al.) | GAN | No | Limited rhythm diversity |
| WaveGAN-ECG (Donahue et al.) | GAN | No | Mode collapse on long signals |
| ECG-VAE (Biswal et al.) | VAE | No | Blurred morphology |
| CoFE-GAN (Jang et al., 2025) | GAN | Yes | Requires iterative latent inversion |
| GCX-ECG (Alcaraz-Segura et al.) | GAN | Yes | Weak morphology preservation |
| Ours | Diffusion | Yes | Single-lead; automated validation only |
Methodology
Pipeline Overview
Our pipeline consists of five stages: data preparation, classifier training, two-stage diffusion model training, counterfactual generation with filtering, and three-regime augmentation evaluation.
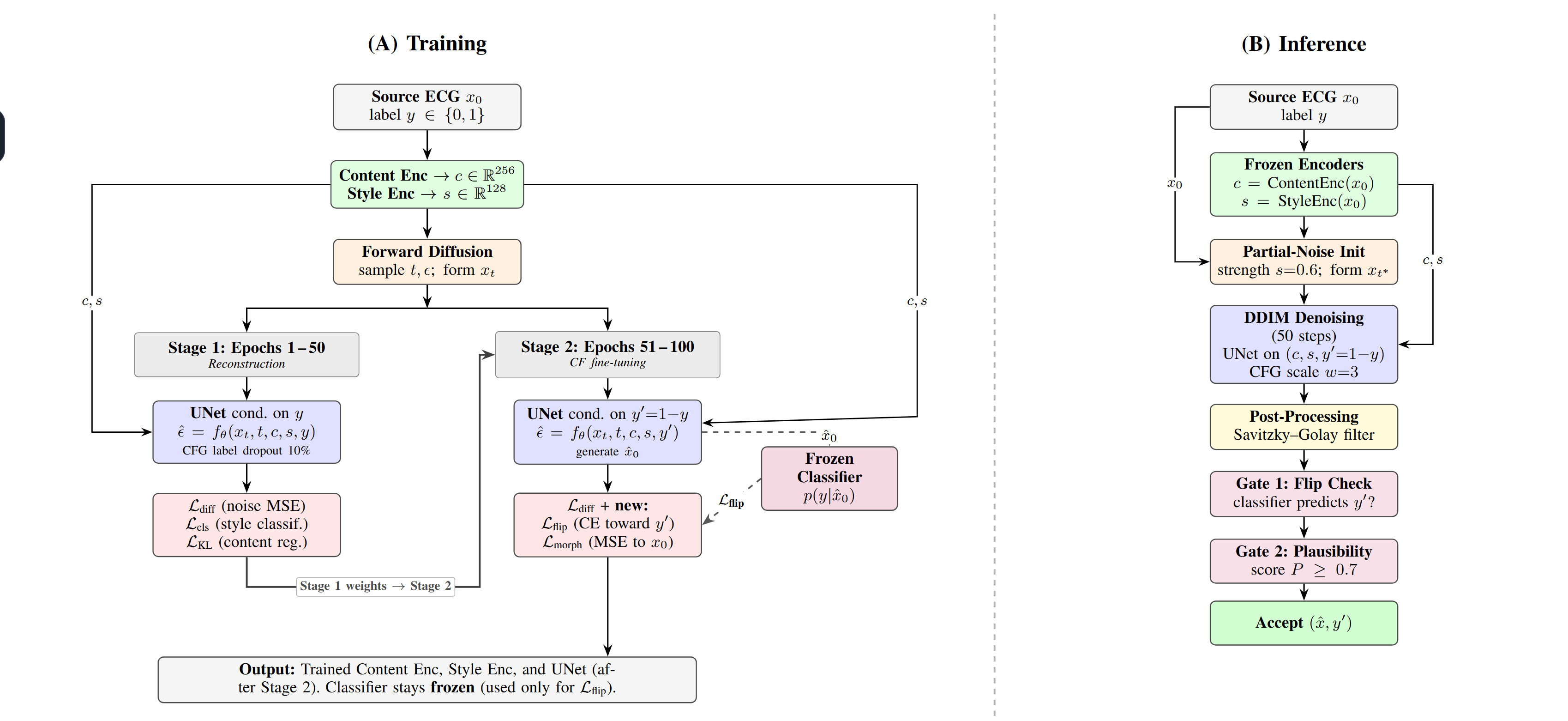
Two-Stage Training
Stage 1 (Epochs 1-50): Reconstruction - The content encoder extracts class-invariant morphological features (256-d VAE latent), and the style encoder captures class-discriminative rhythm features (128-d embedding). The conditional 1D UNet learns to denoise ECG signals with classifier-free guidance (10% label dropout).
Stage 2 (Epochs 51-100): Counterfactual Fine-tuning - The UNet is conditioned on the target class label (y’ = 1 - y), while a frozen AFib classifier provides supervision through a flip loss, and a morphology preservation loss constrains deviation from the source signal.
Inference Pipeline
- Partial-noise initialization - Corrupt source ECG to 60% noise level
- DDIM denoising - 50 steps with classifier-free guidance (scale w = 3)
- Post-processing - Savitzky-Golay smoothing
- Gate 1: Flip verification - Frozen classifier must predict target class
- Gate 2: Plausibility check - Score P = 0.3M + 0.3Phi + 0.4*C >= 0.7
- Accept - Output filtered counterfactual with verified label
Dataset
We use the MIMIC-IV ECG Diagnostic Electrocardiogram Matched Subset (PhysioNet), processing Lead II recordings at 250 Hz, 10-second segments (2,500 samples), bandpass filtered at 0.5-40 Hz.
| Partition | Segments | Normal / AFib |
|---|---|---|
| Training | 104,855 | 52,447 / 52,408 |
| Validation | 22,469 | 11,239 / 11,230 |
| Test | 22,469 | 11,239 / 11,230 |
| Total | 149,793 | 74,925 / 74,868 |
Results
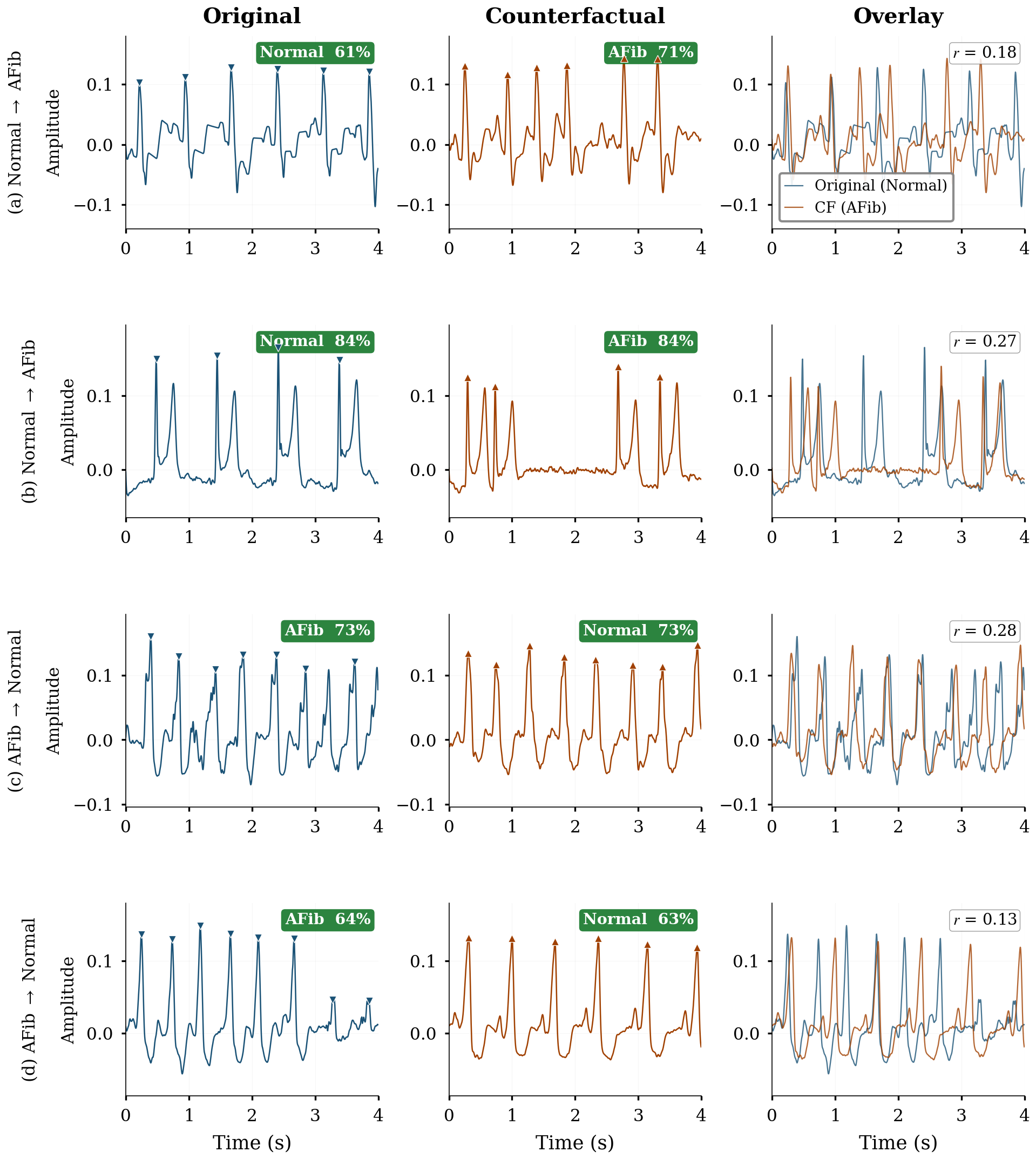
Of 22,469 raw counterfactuals generated, 7,784 passed all quality gates (34.6% acceptance rate).
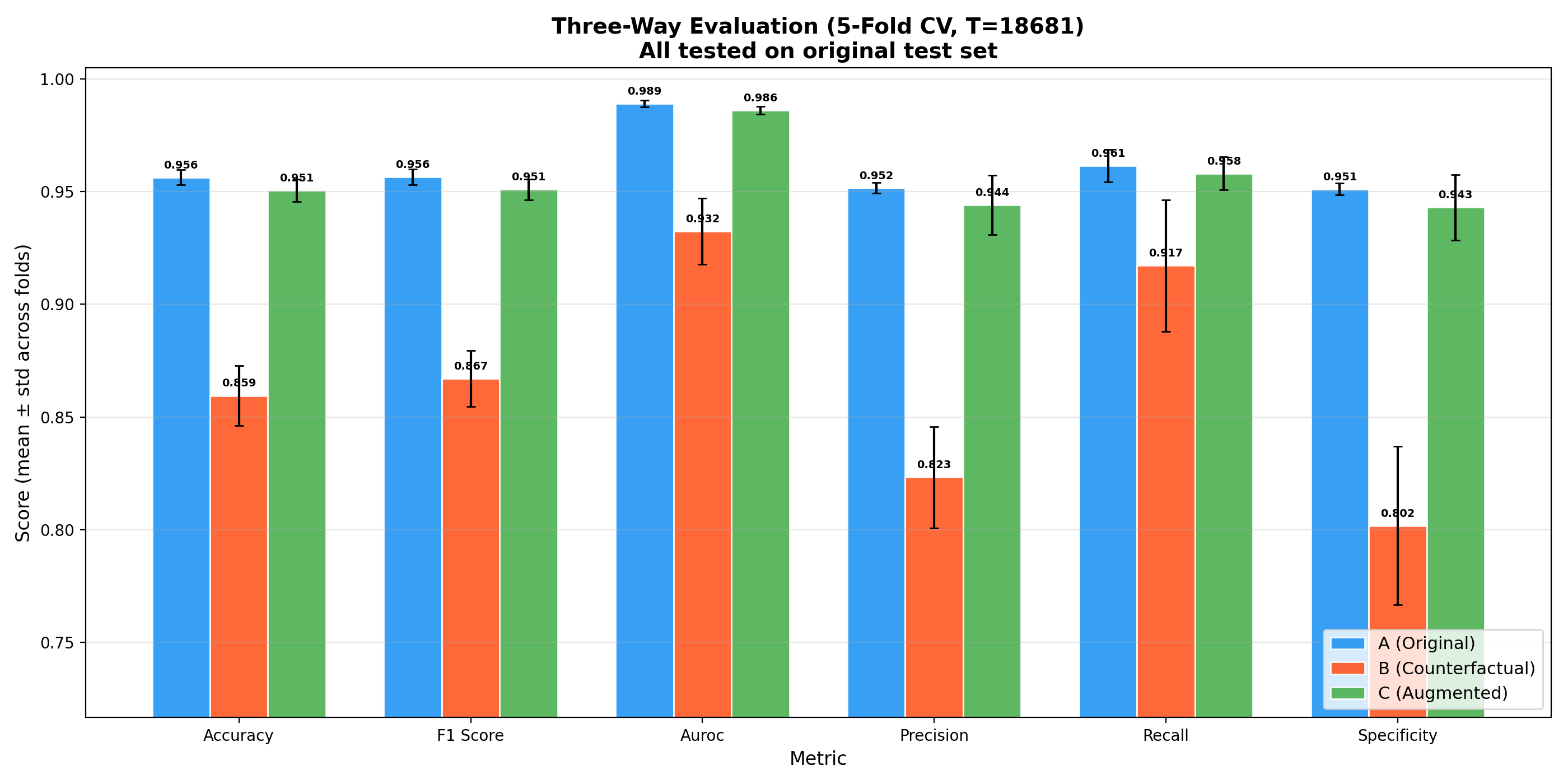
Augmentation Evaluation (5-Fold Cross-Validation)
| Training Regime | Accuracy | F1 Score | AUROC |
|---|---|---|---|
| A - Original only | 95.63 +/- 0.33% | 95.65 +/- 0.35% | 98.90 +/- 0.16% |
| B - CF only | 85.94 +/- 1.32% | 86.70 +/- 1.24% | 93.24 +/- 1.47% |
| C - Augmented (67% + 33%) | 95.05 +/- 0.50% | 95.09 +/- 0.46% | 98.60 +/- 0.17% |
Statistical Equivalence Testing
| Test | Result | p-value |
|---|---|---|
| McNemar’s test | Delta = 0.54% | < 0.001 |
| TOST equivalence (+-2%) | Equivalent | < 0.001 |
| Non-inferiority (2%) | Non-inferior | < 0.001 |
| Dunnett’s (A vs C) | No sig. diff. | 0.340 |
| Dunnett’s (A vs B) | Sig. diff. | < 0.001 |
Conclusion
We presented a diffusion-based counterfactual ECG generation pipeline that transforms single-lead ECG recordings into counterfactuals of the opposing class. The goal is to establish that synthetic counterfactual data can safely substitute for or supplement real patient recordings without degrading diagnostic performance - enabling privacy-preserving data sharing and class balancing for AFib detection.
Publications
- Tharaka Dilshan, Nethmini Karunarathne, Isuri Devindi, Mary M. Maleckar, Jorgen K. Kanters, Roshan Ragel, Isuru Nawinne, Vajira Thambawita. “Diffusion-Based Counterfactual ECG Generation for Atrial Fibrillation Data Augmentation” (2025). Under Review.
Links
- Project Repository
- Project Page
- Model Hub
- Live Demo
- Department of Computer Engineering
- University of Peradeniya
- SEARCH Project (EU Horizon)
This work is part of the European project SEARCH, supported by the Innovative Health Initiative Joint Undertaking (IHI JU) under grant agreement No. 101172997.
